The ultimate goal of our research is to uncover and understand the structural basis of important biological systems and processes, exploiting the unique power of NMR spectroscopy in combination with a range of other biophysical techniques.
Three-dimensional structures of proteins, nucleic acids and their complexes have made enormous contributions to understanding the function of these critically important molecules. With X-ray diffraction (XRD) in the UK in the 1950s, nuclear magnetic resonance (NMR) spectroscopy in Switzerland in the 1970s and later electron microscopy (EM) again in the UK in the 1990s, scientists have developed three different structural biology techniques to determine structures of biomolecules and their complexes over a wide range of molecular sizes and environmental conditions.
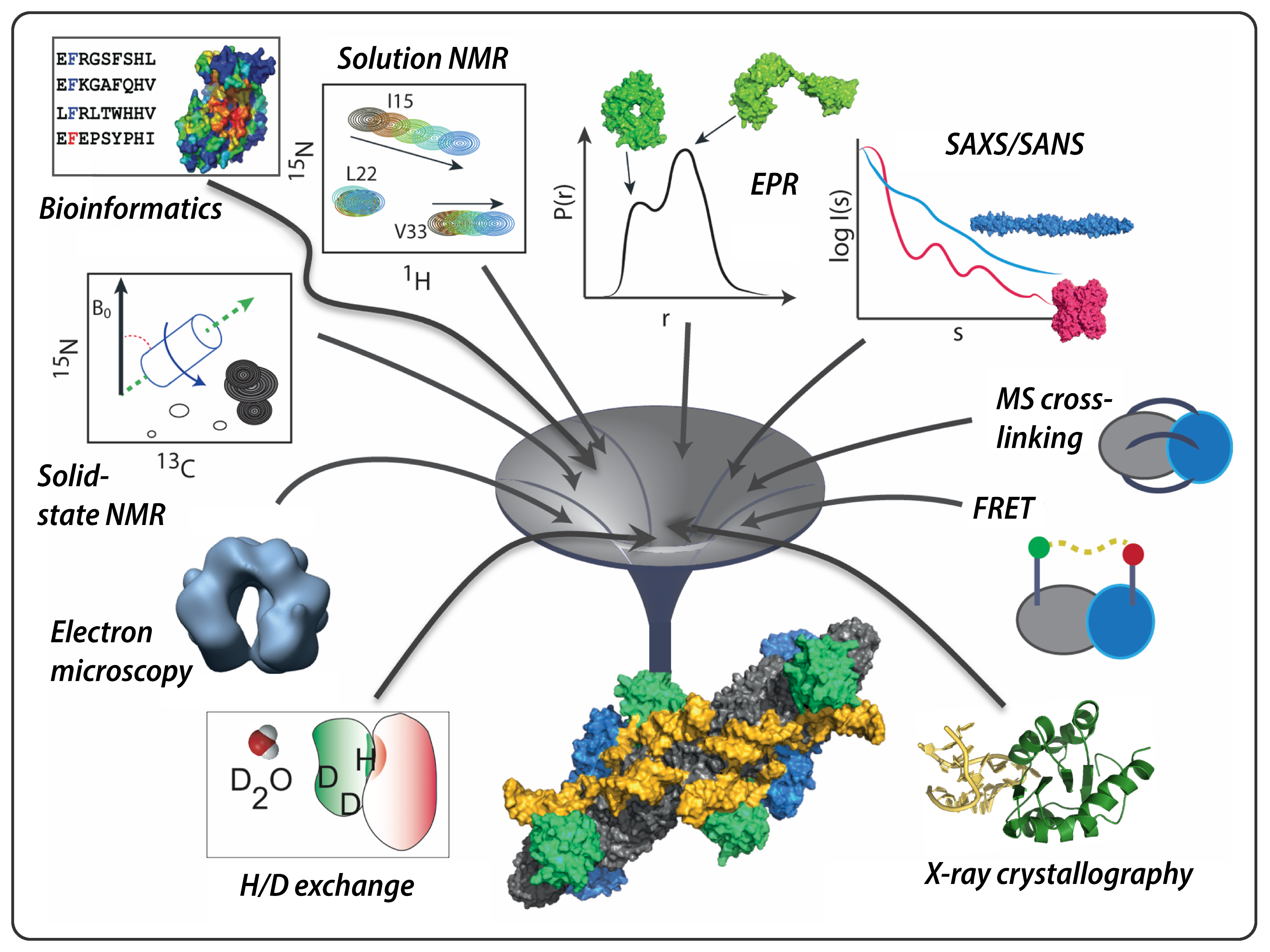
These days, while continuing to solve the structures of as yet uncharacterized systems, we take further steps to investigate the dynamics of biomolecules, ensembles of co-existing conformations, structural changes occurring during function and functional interactions involving disordered biomolecules. When addressing these challenging questions, each of the three techniques encounters limitations if used in isolation. Fortunately, the strengths and weaknesses of XRD, NMR and EM are highly complementary and therefore they can be employed in combination ― together with other lower-resolution structural methods ― in an integrative manner to investigate complex structure–dynamics–function relationships.
In our group, we develop and apply integrative approaches to the structural investigation of biomolecular complexes. We characterize intermolecular interactions at the border between structure, dynamics and disorder by combining NMR spectroscopy (both in solution and in the solid-state) with XRD, EM, small-angle-scattering (SAS) and electron paramagnetic resonance (EPR). In parallel, we develop the computational methodologies necessary to interpret and evaluate the resulting hybrid data (Karaca et al., Nature Methods, 2017).
During the past few years, we have reached some notable milestones; for example, we solved the first structure of a large multi-component enzyme in solution (Lapinaite et al., Nature, 2013), developed protocols to reveal the link between conformational dynamics and function (Graziadei et al., eLife, 2020) and pioneered the application of solid-state NMR (ssNMR) to the structural biology of nucleic acids (Marchanka et al., Angewandte Chemie, 2013; Marchanka et al., Nature Communications, 2015; Ahmed et al., Angewandte Chemie, 2020).
